APO-BrdU™ Kit
View Cart or Continue shopping.
Description
SKU :TNB-6671
The fragmentation of genomic DNA by cellular nucleases during the later stages of apoptosis is also one of the most easily measured features of apoptotic cells. Nuclease activity generates DNA fragments ranging from ~300 bp to 50 bp in length, resulting in a typical DNA ‘laddering’ appearance when analyzed by agarose gel electrophoresis. These fragments have exposed 3’-hydroxyl (OH) ends which can be labeled with bromolated deoxyuridine triphosphates (Br-dUTP). An enzyme, terminal deoxynucleotidyl transferase (TdT), is used to catalyze the template-independent addition of Br-dUTP to the 3’-OH ends of double or single stranded DNA. This method is often called TUNEL (terminal deoxynucleotidyl transferase dUTP nick end labeling) or end labeling. Sites where the Br-dUTP is incorporated can then be detected with an antibody specific to BrdU.
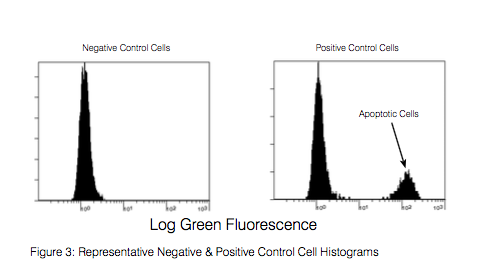
With the APO-BrdU™ Kit cells are first labeled with Br-dUTP, and then sites of incorporation are detected through staining with a FITC anti-BrdU antibody. Samples can then be analyzed via flow cytometry. Samples that are apoptotic will stain brightly with the anti-BrdU antibody due to the substantial number of exposed 3’-OH sites, while cells that are non-apoptotic will not have incorporated significant amounts of Br-dUTP and will stain dimly.
The APO-BrdU Kit is shipped in one container and consists of two packages plus instruction manual. Upon arrival one should be stored at 2-8°C and the other at -20°C.
| Name | APO-BrdU™ Kit |
|---|---|
| Cat. No. | TNB-6671 |
| Clone | PRB-1 |
| Isotype | Mouse IgG1, kappa |
| Format | FITC |
| Application | Flow Cytometry |
