Our Product
High Level Overview
The Cytek Aurora leverages full spectrum technology to provide unprecedented flexibility to scientists, enabling the use of a wide array of new fluorochrome combinations without reconfiguring the system for each application. The Aurora system delivers high-resolution data at the single-cell level to resolve the most challenging cell populations, such as cells with high autofluorescence or low levels of expression of key biomarkers, regardless of assay complexity.
Submit this form to download the paper
Complete the fields below to download the white paper Spectral Analysis Meets Flow Cytometry. Check the consent boxes at the end of the form to receive additional communication from the Cytek Biosciences team.
Learn More About Full Spectrum Cytometry
SpectroFlo® software offers an intuitive workflow from QC to data analysis with technology-enabling tools that simplify running applications.
So Many Colors
Up to 40 colors demonstrated including fluorochromes with emission spectra in close proximity to each other.
Exceptional Sensitivity
Sensitivity redefined using state of the art optics and low noise electronics.
A New Level of Flexibility
Technology
With up to five lasers, three scattering channels, and 64 fluorescence channels, the Aurora system is highly flexible, intuitive and ultra-sensitive. With its intuitive optical design, compact footprint and upgradeability from 3 to 5 laser configurations, the Aurora system suits every laboratory’s needs from simple to high complexity applications.
Proprietary high sensitivity Spectrum Profile Detection Modules enable more efficient spectrum capture for dyes emitting in the 365-829 nm range. Full Spectrum Profiling™ (FSP®) technology captures the full emission signature of every fluorochrome, unlocking its complete expression across the entire light spectrum.
The state-of-the-art optics and low-noise electronics provide excellent sensitivity and resolution. Flat-top beam profiles, combined with a uniquely designed fluidics system, translate to outstanding performance at high sample flow rates.
Testimonials and Publications
Why Cytek® Aurora
Solving Autofluorescence Challenges
Service Plan Comparison
Service Contract Types
Gold
Includes
- Unlimited Coverage Of Parts, Labor & Travel
- 1 Preventative Maintenance Visit/Year
- Priority Response Time
- Unlimited Phone & Email Support
Silver
Includes
- Limited Coverage Of Parts, Labor & Travel
- 1 Preventative Maintenance Visit/Year
- 1 Fully Covered Repair Visit
- Unlimited Phone & Email Support
- 20% Parts Discount
Bronze
Includes
- 1 Preventative Maintenance Visit/Year
- Unlimited Phone & Email Support
- 10% Parts Discount
- All Other Services Billed At Prevailing Rate
Autofluorescence Extraction
The Full Spectrum Profiling technology employed by the Aurora can improve data clarity and resolution with autofluorescence extraction. Certain sample types, such as yeast and tumor samples, present difficulty in resolving due to their highly autofluorescent nature. For applications involving highly autofluorescent particles, the SpectroFlo® CS software autofluorescence extraction tool achieves new levels of resolution and increases sorting performance of challenging samples. SEE POSTER
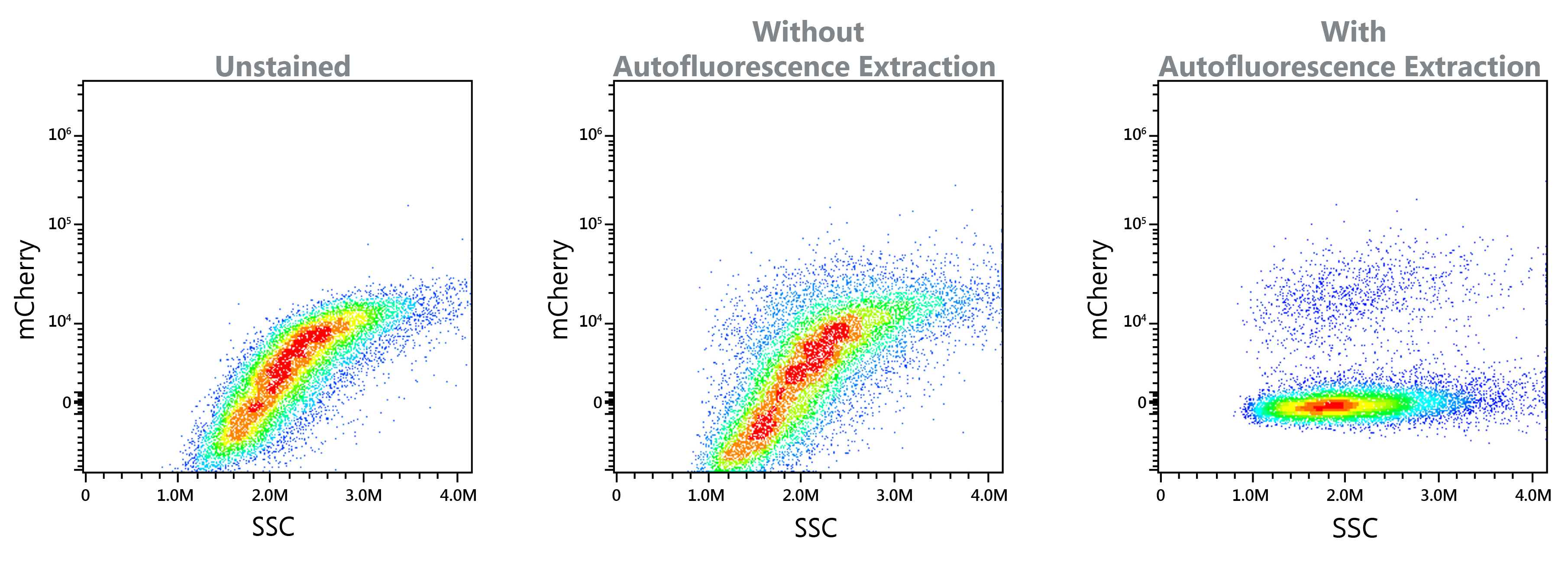
Small Particle Detection
With its onboard 100 mW 405 nm laser and highly sensitive violet SSC detector, Aurora opens the door to a wide variety of small particle applications, taking what was once hidden and placing it in full view.

Submit this form to download the data sheet
Complete the form below to download the "Enhanced Small Particle Detection on Cytek Aurora and Northern Lights" data sheet. Check the consent boxes at the end of the form to receive additional communication from the Cytek Biosciences team.
Deep Immunoprofiling
24 Colors
The three-laser configuration provides outstanding multi-parametric data for a wide array of applications. Check out how markers and fluorochromes in a 24-color panel designed for identification of circulating cell subsets in human peripheral blood look when run on the Aurora system.
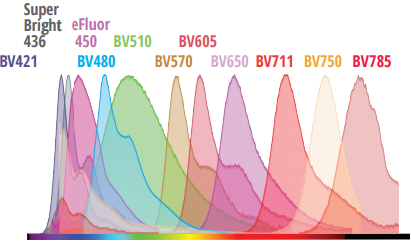
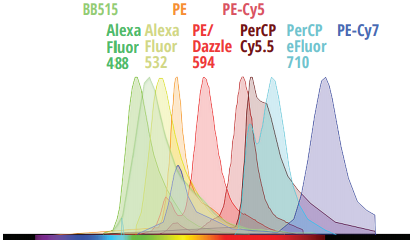
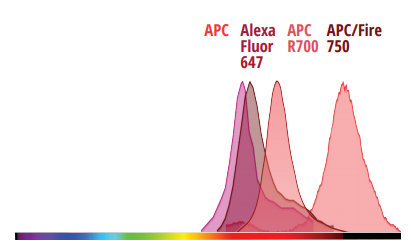
40 Colors
Until recently, developing fluorescence-based flow cytometry assays with 40 colors has been merely aspirational, with many turning to competing technologies for high-parameter applications. No longer. With 64 fluorescence detectors and only 5 lasers, the Cytek Aurora now has the capability to resolve up to 40 colors in combination. Cytek has developed a 40-color human immunophenotyping panel acquired from a single tube, with outstanding resolution.

Class 1 Laser Product.
For Research Use Only. Not for use in diagnostic or therapeutic procedures.

.png)



